Blog with RStudio, R, RMarkdown, Jekyll and Github.
In the first post of this new blog I’ll outline how I’ve set the blog up.
- writing posts in
Rmarkdown - converting posts to markdown from R
- push to Github where Jekyll renders the markdown
- organising all as an RStudio project
What I wanted
I wanted to be able to write about R related things without having to copy and paste code, figures or files. I had used Rmarkdown and knitr before so wanted to use them. I have a wordpress site elsewhere that someone helped me set up a couple of years ago with a blog that I’ve never used. Initially I tried seeing if I could create posts using RMarkdown and put them into that wordpress blog. A brief search revealed that was not straightforward and that Jekyll was the way to go.
What I’ve got
Now I have this blog set up so that I can write all of the posts (including this one) in RMarkdown (.Rmd) and run an R function to convert them to markdown (.md). The blog is hosted for free on Github (you get one free personal site). The site is created using Jekyll on Github, so I didn’t need to install Jekyll or Ruby. I simply edit files locally, then commit and push to Github. I manage the site as an RStudio project, enabling me to edit text, keep track of files and interact with Git all from one interface.
How I got here (steps)
creating Jekyll site on Github
I used Barry Clarks amazing Jekyll-Now repository which you can fork directly from Github and start editing to customize. He gives excellent instructions. What attarcted me to it was that it takes a matter of minutes to set up initially and if you decide you don’t like it you can just delete.
Thanks to Jan Gorecki whose answer on stackoverflow pointed me in this direction and I’ve copied some extra features like the Links and Index pages from his site.
enabling editing of the site from RStudio
I cloned the Github repository for my site using RStudio :
- File, New project, Version control, Clone git
- Repo URL : https://github.com/AndySouth/andysouth.github.io
- Project directory name : andysouth.github.io
setting up so that I can write the posts in RMarkdown
This was the tricky bit for me. I followed inspiration from Jason Bryer and Jon Zelner. I had to tweak them both, the relative paths of figures was my main stumbling block. This was partly because I’m running windows and I couldn’t run the shell scripts that they created. Instead I just run an R function rmd2md which is much the same as Jason’s with some edits to paths and jekyll rendering.
Jason’s function searches a folder that you specify for .Rmd files and then puts .md files into another folder. I set this up so that any plots are put into a third folder. Thus in the root of my site includes these 3 folders.
| Folder | Contents | |
|---|---|---|
| _Rmd | RMarkdown files that I edit | |
| _md | md files created by RMarkdown | |
| figures | plots created by any chunks of R code |
This then means that any R plot is automatically generated, saved as a png and it’s address is written into the md document so that the plot is displayed in the blog. This is shown in a simple example below that queries the WHO API to get the number of cases of one of the forms of sleeping sickness in 2013.
code <- "NTD_4"
year <- 2013
url <- paste0('http://apps.who.int/gho/athena/api/GHO/',code,'.csv?filter=COUNTRY:*;YEAR:',year)
#read query result into dataframe
dF <- read.csv(url,as.is=TRUE)
library(rworldmap)
sPDF <- joinCountryData2Map(dF, nameJoinColumn="COUNTRY", joinCode="ISO3")## 24 codes from your data successfully matched countries in the map
## 0 codes from your data failed to match with a country code in the map
## 220 codes from the map weren't represented in your datamapCountryData(sPDF,nameColumnToPlot="Numeric",catMethod="fixedWidth",mapRegion="africa", mapTitle="Gambian sleeping sickness cases in 2013")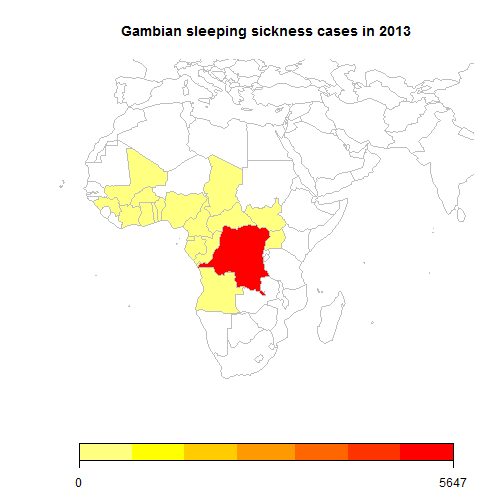
The code syntax highlighting and dark grey background for both code and R text outputs are what come as the default with Jekyll-Now. I’m a little unsure about them. They seem to be specified in _highlights.scss, perhaps I’ll look at modifying later.
If you’d like to look here is the entire source code for the blog and for this individual page.
